For a complete overview of the groups work visit our group repository.
For an overview of select pipelines, please read below.
LotuS2
Updated LotuS pipeline, Dada2 + Unoise clustering, improved speed, LULU and crosstalk postprocessing, offtarget removal, updated tax assignment, better phylogenies and a more rounded user experience.
Available from lotus2.earlham.ac.uk
LotuS
Fast, easy-to-use and highly accurate amplicon sequencing pipeline. Choose between several highly accurate OTU calling algorithms, choose from general or spezialized databases to annotate these taxonomically and create a phylogeny of OTUs - all in on single command.
Available with self-installer from http://psbweb05.psb.ugent.be/lotus/
MG-TK
Pipeline to process metagenomic data from raw reads to taxonomic and functional abudnance matrices. MG-TK offers assembly-dependent (gene catalog) and assembly-independent (direct mapping to appropriate databases) workflows. Previously named “MATAFILER”.
RTK
RTK (rarefaction toolkit) provides fast and scalable rarefaction and diversity estimation for numerical ecology data (e.g. species abundance tables). RTK uses significantly (~100x) less memory while being ~10x faster than competing software. Accessible through R interface - install.package(“rtk”) - or C++ binary:
KSGP
KSGP 16S database for Bacteria, Eukaryotes and Archaeae. Has a significantly higher taxonomic coverage for Archaea and uses GTDB conform taxonomy. Fully integrated with LotuS2.
Database webpage:
AdhesiomeR
AdhesiomeR can be used to classify adhesion related genes in E coli genomes. Available in R or through webserver to directly upload and explore genomes:
https://adhesiomer.quadram.ac.uk/
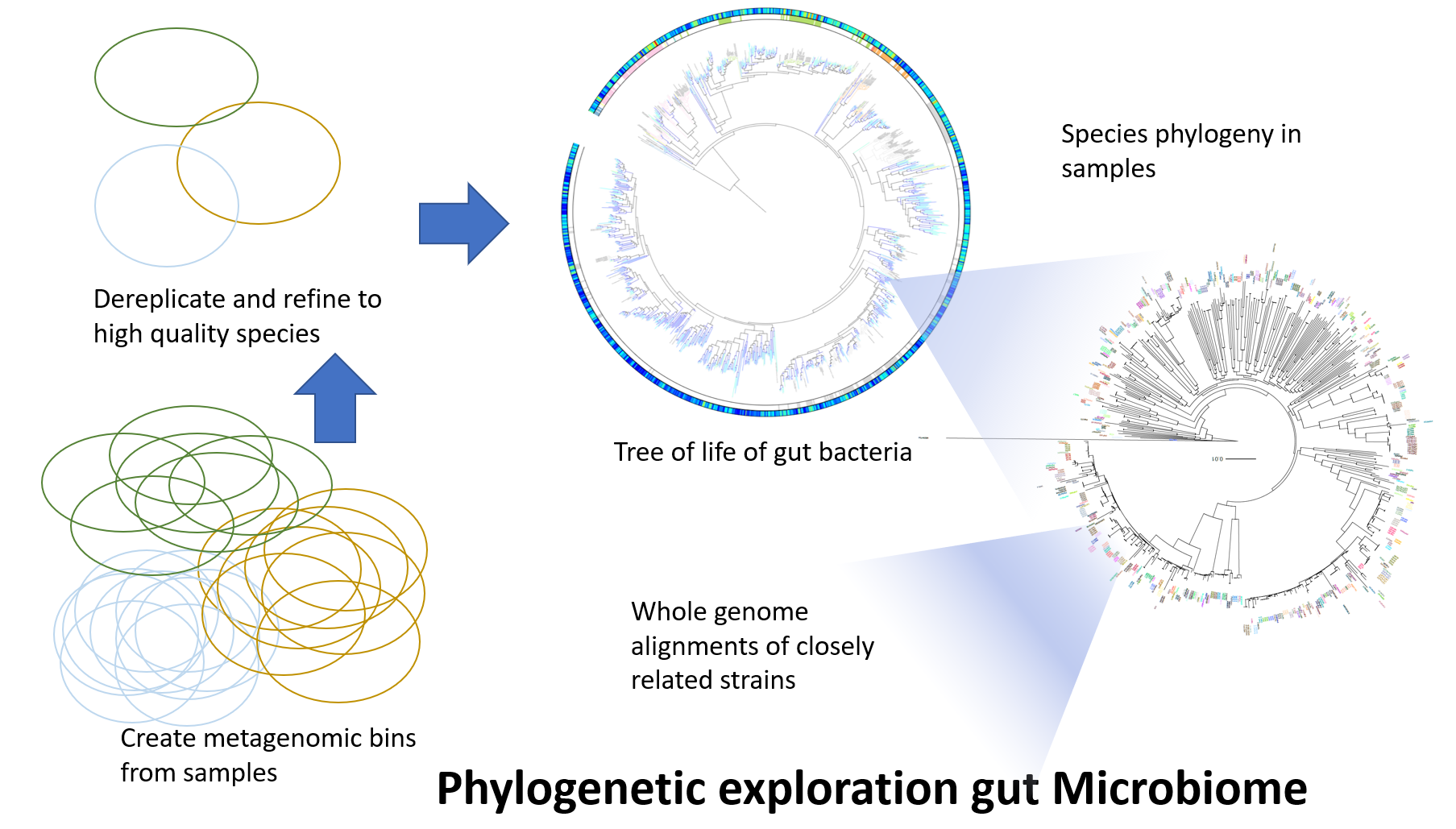
clusterMAGs
An algorithm to cluster MAGs (metagenomic assembled genomes) into metagenomic species (MGS), based on a gene catalogue and universal marker genes. In our benchmarks, this approach is more robust to (biological/artificial) differences in pangenomes and chimeric missassemblies.
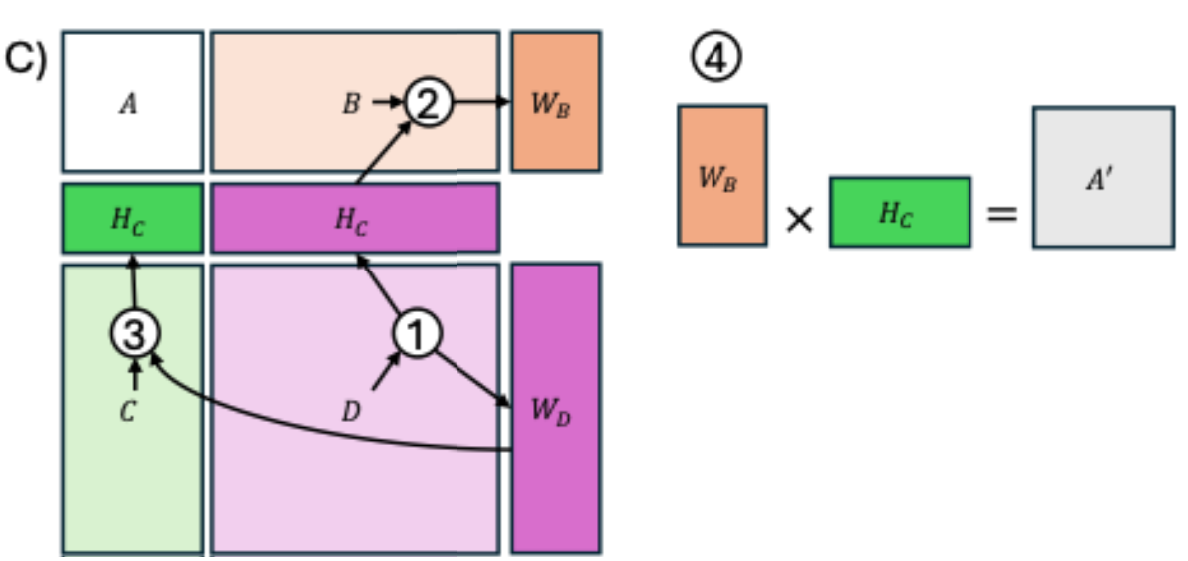
cvaNMF
cvaNMF (chiva-enemef) is an implementation of bicrossvalidation for Non-negative Matrix Factorisation (NMF) rank selection, along with methods for analysis and visualisation of NMF decomposition.
BM-TK
Tool for processing multiple PacBio .bams and predicting methylation status (nextflow).